Surface Functionalization and Patterning by Multifunctional Resorcinarenes by Farid Behboodi-Sadabad
In my latest project, inspired by strong binding ability of natural polyphenols found in plants, I used three different macrocyclic polyphenols, known as resorcin[4]arenes, to modify the surface of a variety of substrates with a uniform and transparent layer by simple dip coating into the dilute solution of these compounds. Eight hydroxyl groups on the large rim of these resorcin[4]arenes provide multiple anchoring points to the surface, while the lower rim decorated with different appending groups introduces desired chemical and physical functionalities to the substrate´s surface. Employing this strategy, I modified the surface with alkenyl functional groups in one step using a resorcinarene compound possessing four alkenyl appending groups on its small rim. Then I used thiol-ene photoclick chemistry to site-selectively post-functionalize the surface with hydrophilic and hydrophobic micropatterns. Thus, resorcin[4]arenes could extend the scope of applications of plant polyphenol and mussel-inspired precursors to tailor-made multifunctional nanocoatings, suitable for a variety of potential applications in biotechnology, biology, and material science.
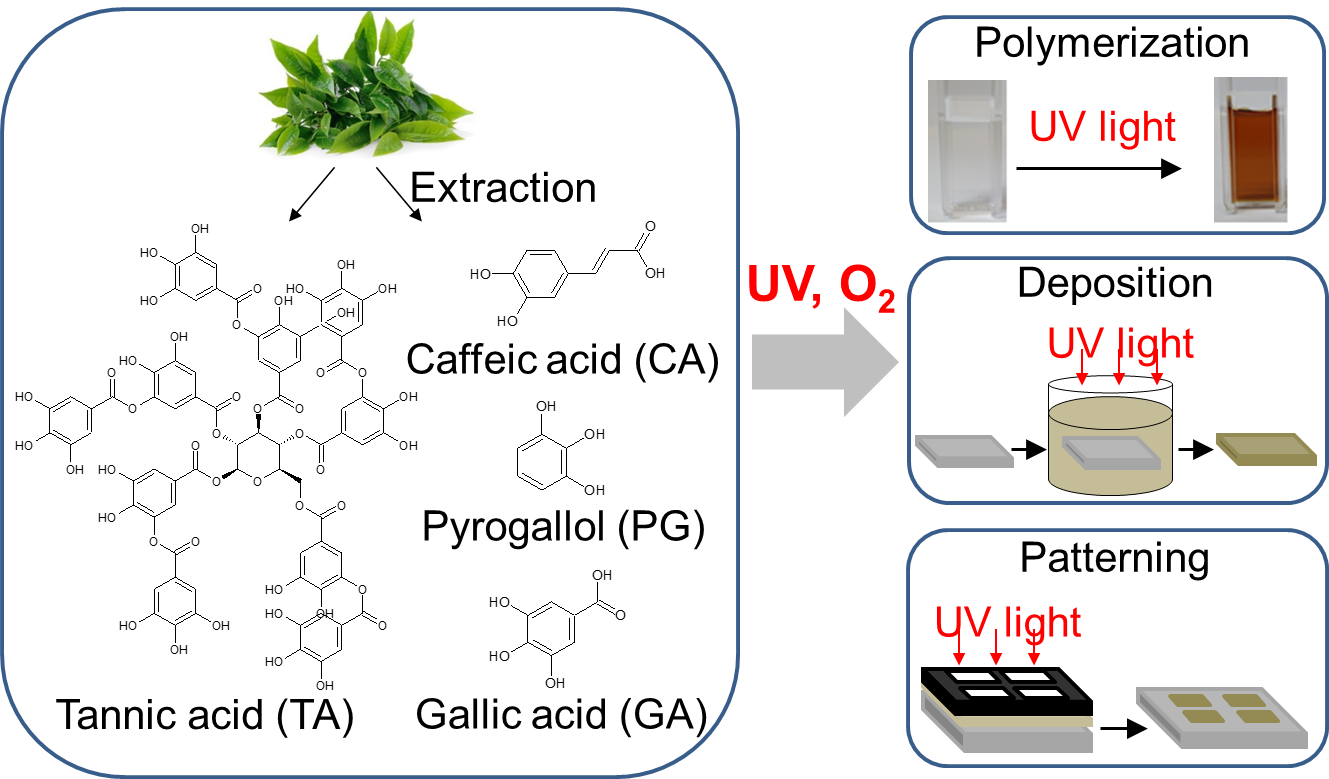
UV-triggered polymerization, deposition and patterning of plant phenolic compounds by Farid Behboodi Sadabad
Nature-inspired strategies have been used extensively to make simple and versatile multifunctional coatings. Plant phenolic compounds can form multifunctional nanocoatings on various substrates, following polymerization under mildly alkaline pH solutions. However, despite a lot of appealing features of such coatings, it is difficult to control this polymerization spatially and temporally, which limits the number of potential applications. I use UV irradiation to trigger polymerization and deposition of these compounds, which opens the possibility to create 2D micropatterns of polyphenol nanocoatings and control this polymerization temporally. This method is general and can be applied to different plant derived phenolic compounds and will be of great interest for health care, anticancer, and drug delivery application.
Reconstruction of 3D Tissue in the Microfluidic µ3DVasc Bioreactor to Investigate Drug Delivery by Eva Zittel
When testing new substances or drugs, the common procedure is an investigation on 2D cell layers. In different assays, cellular uptake, toxicity or specific binding sites are determined. Yet in the next instance, there often is a poor comparability between the results in cell culture experiments to those obtained in the animal model. For example, there are different dose-effect relations or unexpected adverse reactions. A reason for this is that natural surroundings and interactions of cells in an organism (in vivo) do not exist in the culture system (in vitro). For an improved in vitro testing of potential drugs, it is therefore necessary to create a more natural, organotypic micro-environment that allows a 3D arrangement and interactions between different cell types and tissues. Such approaches are summarized as ‘Tissue Engineering’ or ‘Organs-on-a-Chip’. The aim of my project is to develop the co-culture of tissue (like liver or skin) and an already established, microfluidic blood vessel system (µ3DVasc). This allows more natural conditions like a dynamic environment, different cell interactions, introducing drugs via the vascular system or observing a 3D distribution and can therefore improve the reliability of in vitro testing.
NMR Method Development: Robust experiments for the measurement of anisotropic structure parameters by Tony Reinsperger
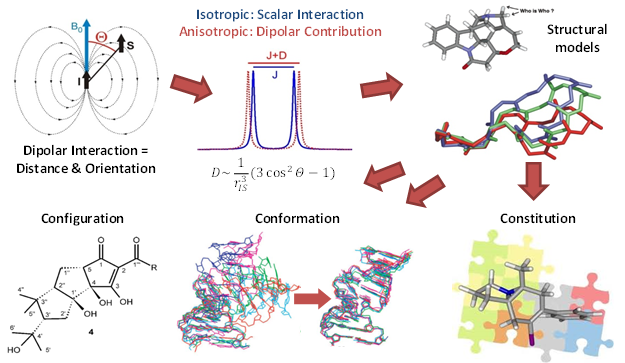
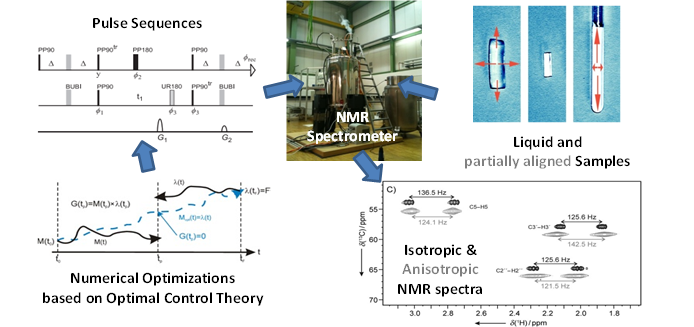 Nuclear Magnetic Resonance (NMR) spectroscopy is a powerful method to study structural aspects of organic materials such as synthetic or natural products and biomolecules in solution with a resolving power down to single atoms. The targets of NMR are the nuclei of chemical elements such as Hydrogen, Carbon and Nitrogen which are highly abundant in living matter. Apart from providing a chemical fingerprint nuclei can interact in the same manner as bar magnets and therefore reveal information about distances and orientations with respect to the external magnetic field.
Nuclear Magnetic Resonance (NMR) spectroscopy is a powerful method to study structural aspects of organic materials such as synthetic or natural products and biomolecules in solution with a resolving power down to single atoms. The targets of NMR are the nuclei of chemical elements such as Hydrogen, Carbon and Nitrogen which are highly abundant in living matter. Apart from providing a chemical fingerprint nuclei can interact in the same manner as bar magnets and therefore reveal information about distances and orientations with respect to the external magnetic field. My work heavily relies on the computer-aided design of NMR experiments by means of numerical optimization based on optimal control algorithms. These robust experiments help to obtain reliable und unambiguous results for structure parameters in order to derive structural models. The refinement of these models is required to answer questions concerning the configuration, conformation and even the basic constitution of molecules.
Functional Block Copolymers as Self-Assembling Units for Synthetic Enzyme Cascades by Hatice Turgut
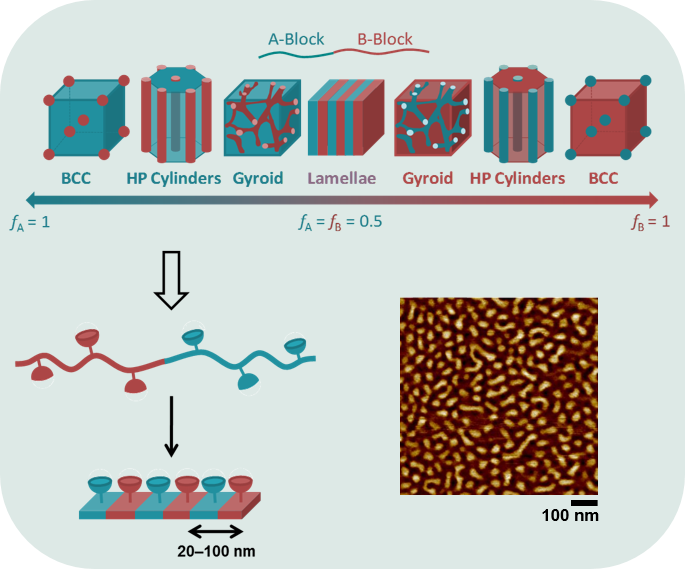
Block copolymers have gained popularity in many areas such as nanolithography, drug delivery and nanoreactors. When they are spin-coated to form a thin film on a solid substrate and depending on the volumic fractions of the blocks, they can phase-separate and self-assemble into nanosized motifs. We exploit these motifs to spatially order molecules, particularly enzymes. Upon immobilization, enzymes can for instance acquire enhanced stability, possibility for repeated and continuous use, or modulation of catalytic activity. In my PhD project, I synthesize functional block copolymers to create nanostructured reactive surfaces in order to attach multiple enzymes with a close-to-single-enzyme resolution. Modern polymer chemistry protocols, bionconjugation methods, and surface characterization techniques such as atomic force microscopy are my daily tools.
TAL effector mediated regulation of regulatory elements in zebrafish by Kathrin Herbst
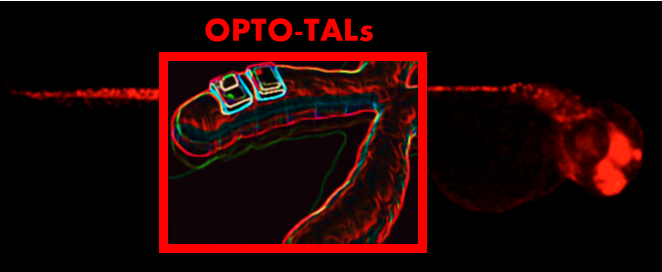

Providing a key intermediate model between super high throughput in vitro screens and the classic mammalian models, zebrafish combines the accessibility of in vitro models and the complex functional capabilities of mammalian models. The completed genome provides us with a roadmap to study the function of genes and together with the rapid emergence of genome editing tools it provides us with the opportunity to unravel the networks underlying molecular mechanisms in vertebrates. To study these processes in vivo, we established the TAL tailoring technique as a molecular tool to control zebrafish gene expression by the help of customized TAL activators and repressors. To enable the control of those TAL effectors in a temporally manner we are furthermore (in collaboration with H. Beyer (AG Weber; Freiburg)) developing an optogenetic two-hybrid system integrating the customizable TALE DNA-binding domain of Xanthomonas sp. with a light-sensitive expression system of Arabidopsis thaliana. With that system we will establish a novel mode of optogenetic control which enables us to study the role of endogenous transcriptional regulatory elements in zebrafish.
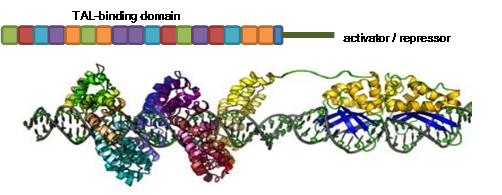
Luminescent copper complexes by Manuela Wallesch
I am working on luminescent copper complexes, which are used as labels for biological systems and emitters for organic light emitting diodes. These complexes are favorable for applications in material science as they exhibit high luminescence quantum yields and are especially stable under excitation. To develop a better understanding of the detailed properties of the synthesized complexes I am using x-ray spectroscopic techniques to study their detailed electronic and geometrical structure. This will help us to further improve and modify the luminescent copper complexes specifically.
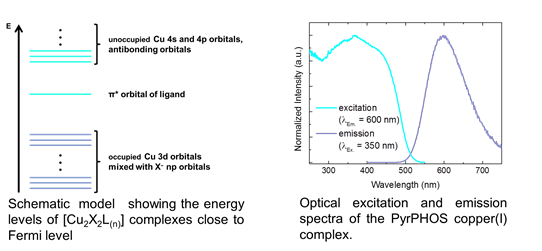
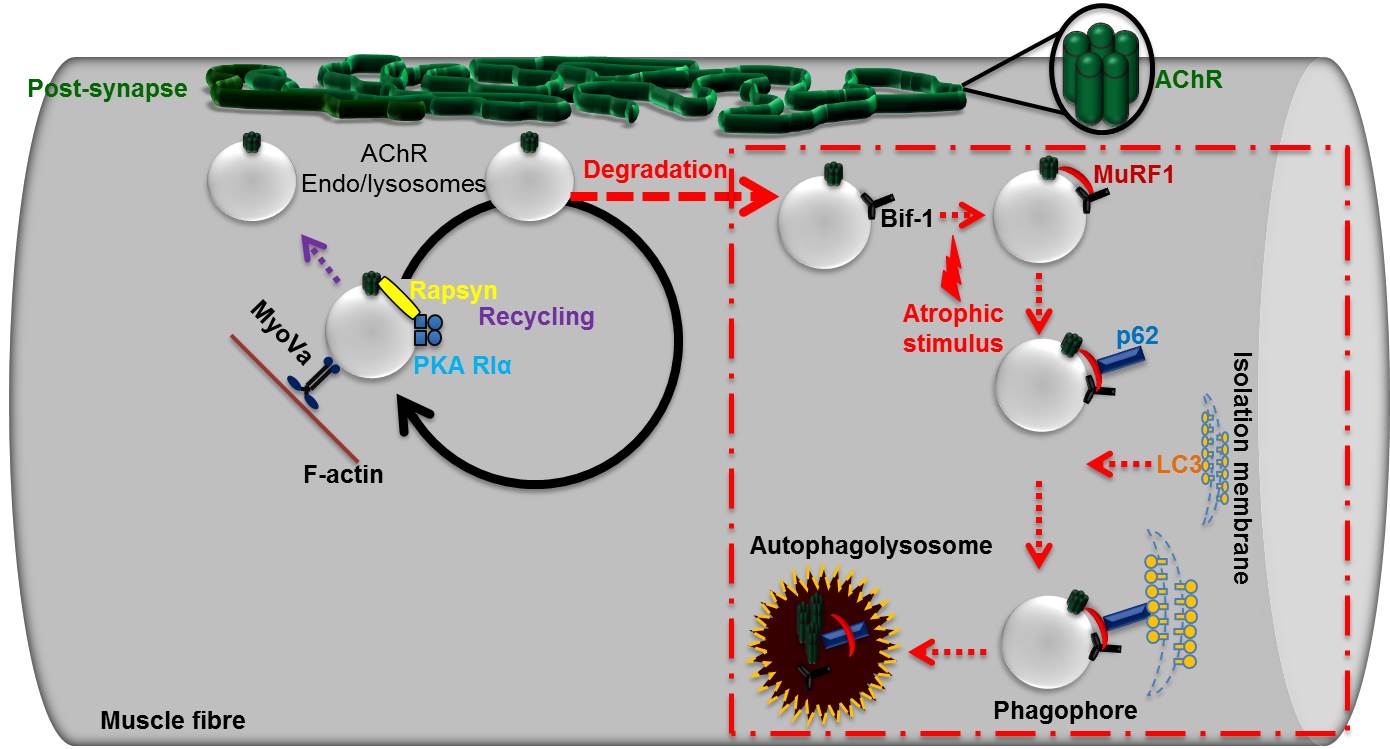
Muscle and muscle contraction- understanding on a molecular level by Muzamil Khan
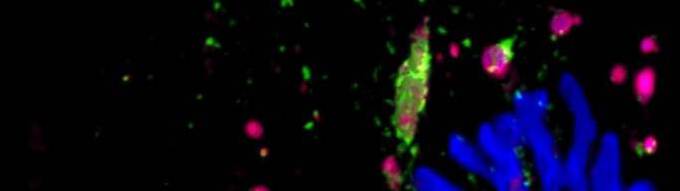
Vertebrate neuromuscular junction is the connection (synapse) between a motor neuron and skeletal muscle fibre. The acetylcholine (Ach) released from the motor neuron binds to the Ach receptors present on the muscle surface, the binding of ACh to AChR evokes skeletal muscle fibre contraction. Ailments affecting AChRs can be very detrimental for the health of the body. AChRs are increasingly degraded upon various pathophysiological conditions such myasthenic syndrome, loss of muscle mass (muscle atrophy), aging. Using live microscopic technique, our group's studies have elucidated various molecular partners of AChR trafficking (Figure). Currently we are studying how AChRs are degraded upon muscle atrophying conditions (red square on the right side).